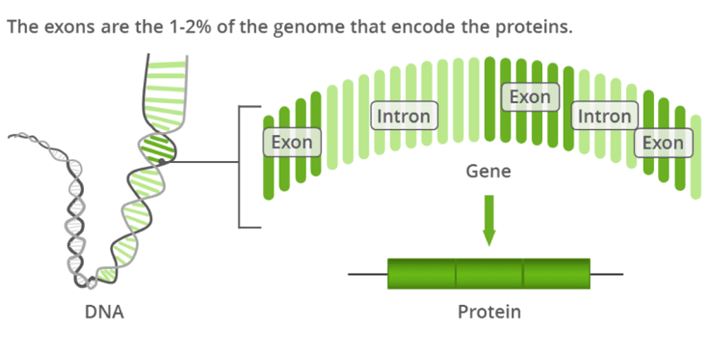
Whole exome sequencing (WES) is one of the most popular next-generation sequencing (NGS) methods in use today for the detection of genetic variants. WES uses specific technology to capture approximately 1% of the entire human genome (the human exome).
What is whole exome sequencing?
Whole-exome sequencing refers to the capture of the coding region (exonic region) of the genome. This method focuses on the protein-coding region of DNA, which makes up approximately 1% of the entire genome (1% of millions of DNA "letters").
Exomes are sequenced because they contain many of the genetic variants associated with common diseases. These include single nucleotide polymorphisms (SNP), variation in the number of copies (CNV), structural variants (SV) and detection of SNV (single nucleotide variants).
In addition, exome sequencing allows us to identify rare variants, which are less frequent than SNPs but occur much more frequently than CNVs and SVs. Rare variants are responsible for Mendelian disorders (genetic diseases), some cancers and complex diseases like diabetes and autism.
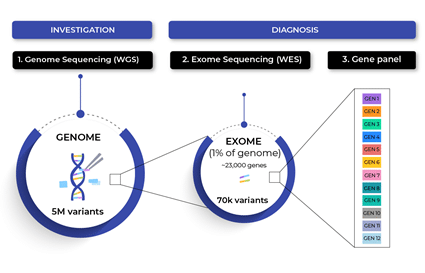
What is the difference between whole exome sequencing (WES) and whole genome sequencing (WGS)?
Whole Genome Sequencing sequences the entire genome (DNA samples are required) to provide a full picture of the entire human genome. In contrast to whole-genome sequencing, WES involves sequencing only the protein-coding portion of the genome (the exons) allowing to exclusively detect exome variants.
Whole exome sequencing is an important powerful tool to identify disease-causing mutations. This method allows researchers to identify the underlying cause of a patient's symptoms without having to sequence every gene in his or her body. However, it has some limitation. While whole genome sequencing is effective in detecting large deletions, duplications and chromosomal rearrangements, WES cannot detect these types of alterations.
However, both technologies are valid for clinical diagnostics.
How does whole exome sequencing work?
The process begins with DNA sample extraction. This step isolates the genetic material from cells, including both nuclear and mitochondrial DNA. Next, it uses PCR technology to amplify the extracted DNA into millions of copies. After purifying and concentrating the amplified product, it attaches adapters to each end of the fragments. These adapters help prepare the DNA for sequencing by adding indexes that identify the specific fragment being sequenced. Finally, the purified DNA is loaded and the exonic regions undergoes massively parallel sequencing.
What are the benefits of whole exome sequencing?
Whole-exome sequencing is a less costly way of identifying genetic variations associated with genetic diseases. Exomes are particularly useful for individuals whose family history suggests a strong likelihood of inherited diseases.

The process involves isolating the DNA from a patient sample and preparing it for next-generation sequencing. Next, researchers use high throughput technologies to sequence each individual base pair within the targeted gene region. The resulting information is used to identify mutations that are linked to specific disorders.
- In cases where the causative mutation for a mendelian disease is known, WES can be used to identify additional mutations in the same gene (e.g., CFTR gene for cystic fibrosis). This helps to determine whether there is one dominant mutation or several recessive ones.
- In cases where the cause of the disease is still unknown, WES allows us to look for novel mutations in known candidate genes.
- In some polygenic diseases, such as autism spectrum clinical diagnosis, it is likely that many different alleles contribute to the risk for developing the disease. In these situations, WES provides a powerful tool to investigate the role of each allele in causing the disease.
Comparison with other approaches
Exome sequencing is particularly useful in identifying the genetic basis and pathogenic mutations of Mendelian diseases, where there is usually a single mutation causing the disorder. This approach is much more efficient than traditional methods of diagnosis, which require testing each gene individually.
This technique is also becoming increasingly popular for use in cancer research, since many cancers arise due to somatic changes in specific genes rather than inherited germline mutations.
Complete exome price
When we talk about cost of sequencing, WES is more affordable than WGS as it is looking into fewer regions of the DNA although it is still a challenge. There are other techniques that allow us to sequentially analyze small parts of the genome at reduced cost such as el genotyping. These methods make it easier to obtain an understanding of our genetic code.
Take your knowledge to the next level with ADNTRO. Choose between our genotyping service or whole genome sequencing and start making decisions based on your unique science.