In ADNTRO, the first step to begin your journey through your DNA is to take a saliva sample. The shows the we send to analyze Eurofins Lab, in Denmarkto, one of the largest genotyping centers in europe. The laboratory analyzes the sample using the technique Global Screening Arrayand provides us with the data that we process at ADNTRO in order to be able to provideyou all the content that you will see in the demo of our website.
How does the laboratory analyze the saliva sample?
In the sample are cells of the wall of the mouth that were washed away by saliva. These cells contain the DNA that will be analyzed by the technique of Global Screening Array.
First, the DNA is extracted through a procedure that breaks the cell membranes and separates the internal components, leaving only the Purified DNA. This DNA will be cloned thousands of times using the technique PCR and replacing the nucelotides natural for a few fluorescently labeled.
Then the DNA will be fragmented into small pieces or regions. These regions include specific genes that will allow us to find out, depending on what SNPs (variants of a nucleotide) have, possible phenotypic characteristics such as eye color, a food intolerance or a specific ethnic origin.
What is a DNA chip in a Global Screening Array?
On the other hand, a solid surface of about 2 x 2 cm called DNA chip. The chip contains 700,000 positions or wells in which a DNA probe for each specific SNP. This is a DNA sequence that corresponds to the part of a gene in which a specific SNP is found. On this chip, with the help of automatic devices capable of depositing very small drops in defined positions, our cloned and fragmented DNA sample is poured and allowed to incubate long enough for the DNA can hybridize.

If the sample contains the SNP variant that matches the probe on the chip, the DNA strands will hybridize. In this way, the sample DNA fragment is bound to the probe. As the probe is fixed to the plate, we can wash the chip and the sample fragment will not be removed, unless it does not contain the SNP variant of the probe, in which case it will be eliminated with the wash as it has not hybridized.
After incubation and washing, It's analized the chip with a machine capable of recognizing fluorescence in all positions. Those that contain the hybridized sample DNA fragment will be those that have fluorescence while those that have not hybridized, the fragment will have been removed and with it the fluorescence.
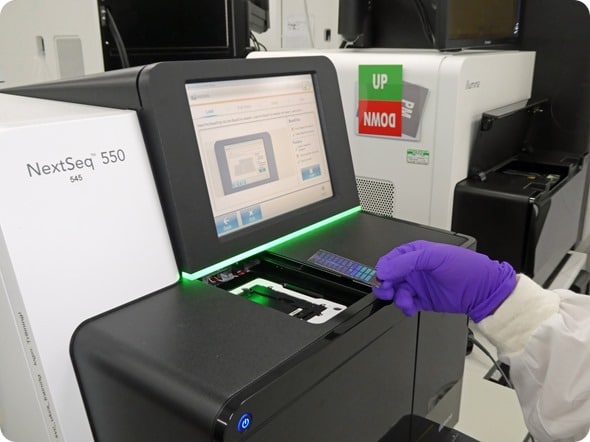

With this, the Global Screening Array ends, obtaining all the raw data that Eurofins will send us for analysis in ADNTRO. The only thing left is that explore your own DNA in your ADNTRO personal space.
Buy your DNA test or upload your RAW DNA data!