A history of population genetic variation.
The title gives name to an idea developed by ADNTRO and that has two important objectives; first help you understand how different your genotype is from the population mean (in this case using 1000 genotypes with which to compare - with a mixture of African, Asian and European populations to a greater extent) and on the other hand to identify some genotypes that are currently under positive selection.
Explaining, in detail, the genetic analysis carried out is not the objective of this article, but it is worth establishing some basic fundamentals.
The 1000 Genome project (The 1000 Genomes Project Consortium 2010) has identified 15 million common DNA variants, mostly single nucleotide polymorphisms (SNPs), constituting a key tool for investigations of the extent of genetic variation in human populations, and is the basis of the study. Figure #1 shows how these variations are shared by different populations:
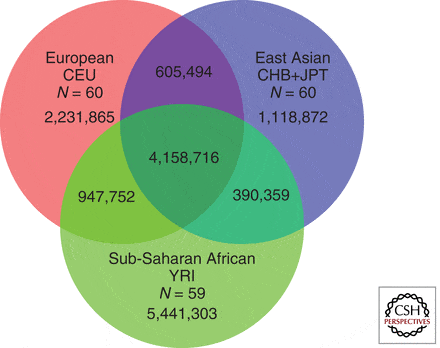
SOURCE: COLD SPRINGS LAB
The 1000 Genomes Project has shown, for example, that populations of African descent have the highest proportion of new variants, reflecting the greater diversity of African populations. However, as Asian and European populations have expanded more rapidly in the recent past, it has been suggested that sequencing a greater number of individuals from each population could produce a greater number of polymorphic sites in Eurasians than in Africans (Gravel et al. 2011).
All data sets have provided information on the degree of population genetic makeup and levels of mixing in many populations around the world.
If 1000G is the basis for our study on How different am I from the 1000G average… and how is that average obtained?
This last point is very important, because when we compare your DNA with a "population mean" what we are actually doing is applying the equation of Hardy-Weinberg explained by Khan Academy in that short video, with an example about recessive / dominant genes like eye color. - This model is very simplistic, but it allows to obtain a starting point with which to compare, assuming that the population is in genetic equilibrium; which is false and would take us to the second point about positive selection and the Neutral Model.
Based largely on a brilliant series of articles by Kimura in the 1960s and '70s, the neutral model of evolution has become the standard against which positive selection must be detected. In the neutral model, the vast majority of mutations are divided into two groups. The first group, for which the model is named, are selectively neutral (or near neutral) mutations that fixate on a species by genetic drift. These changes explain almost all the observable nucleotide changes between two species.
The second group are selective mutations, which continually arise and are eliminated over time by natural selection. Because these mutations are eventually eliminated from one species, they are rarely observed when comparing the genomes of two species. On the other hand, they are the basis for a substantial fraction of the population diversity within a species. Because they cause mutant phenotypes, these mutations are well known to functional geneticists, since they represent almost all mutant strains and human diseases that are widely studied throughout human biology and health - We still have a lot to learn ... but some examples that are encompassed within this positive selection are well known:
- Hemophilia (predominant in Africa because it protects against malaria...but which is gradually disappearing from our DNA) - and a serious health condition since red blood cells carry significantly less oxygen than in people without this condition.
- The Lactose intolerance, very predominant in Asian populations, but on the contrary - in its opposite version, that is, lactose tolerant - predominant in European populations, due to generations fed with foods rich in milk
- The eye color blue; a recessive mutation, that is, you must inherit both alleles from father and mother to have blue eyes - but one that has not been eliminated from our DNA (if not even the frequency is believed to have increased) due, potentially, to a ' improvement »in the success of finding a partner.) -. All this are scientific hypotheses not yet proven, but the evidence exists (increase in allelic frequency)
Finally, to make it easier to understand, in the results of the DNA testing from ADNTRO We played with the data to provide you with a relative measure of your "level of differentiation" from this average, based on standard deviations, which we explain in more detail in your results.